-Search query
-Search result
Showing all 50 items for (author: hau & jl)

EMDB-29281: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-29282: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-15089: 
Sodium pumping NADH-quinone oxidoreductase with substrates NADH and Q2
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-15088: 
Sodium pumping NADH-quinone oxidoreductase
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-15090: 
Sodium pumping NADH-quinone oxidoreductase with substrate Q2
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-15091: 
Sodium pumping NADH-quinone oxidoreductase with substrate Q1
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-15092: 
Sodium pumping NADH-quinone oxidoreductase with inhibitor DQA
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-15093: 
Sodium pumping NADH-quinone oxidoreductase with inhibitor HQNO
Method: single particle / : Hau JL, Kaltwasser S, Vonck J, Fritz G, Steuber J

EMDB-29686: 
N2 neuraminidase of A/Tanzania/205/2010 H3N2 in complex with 4 FNI19 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29704: 
N2 neuraminidase of A/Tanzania/205/2010 H3N2 in complex with 3 FNI9 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29705: 
N2 neuraminidase of A/Tanzania/205/2010 H3N2 in complex with 4 FNI9 Fab molecules
Method: single particle / : Dang H, Snell G

EMDB-29706: 
N2 neuraminidase of A/Hong_Kong/2671/2019 in complex with 3 FNI9 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29707: 
N2 neuraminidase of A/Hong_Kong/2671/2019 in complex with 4 FNI9 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29708: 
N2 neuraminidase of A/Tanzania/205/2010 H3N2 in complex with 3 FNI17 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29709: 
N2 neuraminidase of A/Tanzania/205/2010 H3N2 with S245N S247T mutations in complex with one FNI17 Fab molecule
Method: single particle / : Dang HV, Snell G

EMDB-29710: 
N2 neuraminidase of A/Hong_Kong/2671/2019 in complex with 4 FNI19 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29711: 
Neuraminidase of B/Massachusetts/02/2012 (Yamagata) in complex with 4 FNI17 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-29712: 
N2 neuraminidase of A/Hong_Kong/2671/2019 in complex with 3 FNI19 Fab molecules
Method: single particle / : Dang HV, Snell G

EMDB-27036: 
N332-GT2 trimer in complex with pooled polyclonal fab from BG18-exposed mice (day 14)
Method: single particle / : Jackson AM, Ward AB

EMDB-27037: 
N332-GT2 trimer in complex with pooled polyclonal fab from wild type mice (day 42)
Method: single particle / : Jackson AM, Torres JL, Ward AB

EMDB-27038: 
N332-GT2 trimer in complex with pooled polyclonal fab from BG18-exposed mice (day 42)
Method: single particle / : Jackson AM, Nogal B, Ward AB

EMDB-27039: 
SARS-CoV-2 S protein mix in complex with Novavax NHP 36057 polyclonal Fab
Method: single particle / : Jackson AM, Bangaru S, Ward AB

EMDB-27040: 
SARS-CoV-2 S protein mix in complex with Novavax NHP 36964 polyclonal Fab
Method: single particle / : Jackson AM, Bangaru S, Ward AB

EMDB-27041: 
SARS-CoV-2 S protein mix in complex with Novavax NHP 37452 polyclonal Fab
Method: single particle / : Jackson AM, Bangaru S, Ward AB

EMDB-27042: 
SARS-CoV-2 S protein mix in complex with Novavax NHP 37611 polyclonal Fab
Method: single particle / : Jackson AM, Bangaru S, Ward AB

EMDB-27197: 
SARS-CoV-2 spike protein complexed with polyclonal Fab from donor D (Donor Lotus-25)
Method: single particle / : Bangaru S, Jackson AM, Ward AB

EMDB-27198: 
SARS-CoV-2 spike protein complexed with polyclonal Fab from donor C (Donor 1992)
Method: single particle / : Bangaru S, Sewall LM, Ward AB

EMDB-27199: 
SARS-CoV-2 spike protein complexed with polyclonal Fab from donor B (Donor 1989)
Method: single particle / : Bangaru S, Sewall LM, Ward AB

EMDB-27200: 
SARS-CoV-2 spike protein complexed with polyclonal Fab from donor A (Donor 1988)
Method: single particle / : Bangaru S, Sewall LM, Ward AB

EMDB-24109: 
Inactivated state of 2-APB-bound wildtype rat TRPV2 in nanodiscs
Method: single particle / : Pumroy RA, Protopopova AD, Gallo PN, Moiseenkova-Bell VY

EMDB-24110: 
Activated state of 2-APB-bound wildtype rat TRPV2 in nanodiscs
Method: single particle / : Pumroy RA, Protopopova AD, Gallo PN, Moiseenkova-Bell VY

EMDB-25650: 
Activated state of 2-APB and CBD-bound wildtype rat TRPV2 in nanodiscs
Method: single particle / : Pumroy RA, Protopopova AD

EMDB-25651: 
Inactivated state of 2-APB and CBD-bound wildtype rat TRPV2 in nanodiscs
Method: single particle / : Pumroy RA, Protopopova AD

EMDB-13847: 
Cryo-EM structure of native human A2ML1
Method: single particle / : Zarantonello A, Nielsen NS, Andersen GR

EMDB-13848: 
Structure of TEV cleaved A2ML1 (A2ML1-TE)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

EMDB-13849: 
Structure of TEV conjugated A2ML1 (A2ML1-TC)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

EMDB-13850: 
Structure of TEV cleaved A2ML1 dimer (A2ML1-TT dimer)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

PDB-7q5z: 
Cryo-EM structure of native human A2ML1
Method: single particle / : Zarantonello A, Nielsen NS, Andersen GR

PDB-7q60: 
Structure of TEV cleaved A2ML1 (A2ML1-TE)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

PDB-7q61: 
Structure of TEV conjugated A2ML1 (A2ML1-TC)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

PDB-7q62: 
Structure of TEV cleaved A2ML1 dimer (A2ML1-TT dimer)
Method: single particle / : Nielsen NS, Zarantonello A, Andersen GR

EMDB-7321: 
Integrative Structure and Functional Anatomy of a Nuclear Pore Complex
Method: subtomogram averaging / : Kim SJ, Fernandez-Martinez J, Nudelman I, Shi Y, Zhang W, Ludtke SJ, Akey CW, Chait BT, Sali A, Rout MP

EMDB-5455: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with soluble 2-domain CD4
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5456: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with 17b IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5457: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC01 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5458: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC03 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S
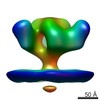
EMDB-5459: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC02 Fab
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5460: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC02 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S
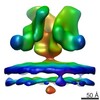
EMDB-5461: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC01 IgG and 17b IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5462: 
Cryo-electron tomography of a trimeric soluble HIV Env construct, gp140 SOSIP, in complex with Fab 17b
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
